The joy of algorithms and NoSQL revisited: the MongoDB Aggregation Framework
Part 1 of this article describes the use of MongoDB to implement the computation of molecular similarities. Part 2 discusses the refactoring of this solution by making use of MongoDB’s build-in map-reduce functionality to improve overall performance. Part 3 finally, illustrates the use of the new MongoDB Aggregation Framework, which boosts performance beyond the capabilities of the map-reduce implementation.
[/information]In part 1 of this article, I described the use of MongoDB to solve a specific Chemoinformatics problem, namely the computation of molecular similarities through Tanimoto coefficients. When employing a low target Tanimoto coefficient however, the number of returned compounds increases exponentially, resulting in a noticeable data transfer overhead. To circumvent this problem, part 2 of this article describes the use of MongoDB’s build-in map-reduce functionality to perform the Tanimoto coefficient calculation local to where the compound data is stored. Unfortunately, the execution of these map-reduce algorithms through Javascript is rather slow and a performance improvement can only be achieved when multiple shards are employed within the same MongoDB cluster.
Recently, MongoDB introduced its new Aggregation Framework. This framework provides a more simple solution to calculating aggregate values instead of relying upon the powerful map-reduce constructs. With just a few simple primitives, it allows you to compute, group, reshape and project documents that are contained within a certain MongoDB collection. The remainder of this article describes the refactoring of the map-reduce algorithm to make optimal use of the new MongoDB Aggregation Framework. The complete source code can be found on the Datablend public GitHub repository.
1. MongoDB Aggregation Framework
The MongoDB Aggregation Framework draws on the well-known linux pipeline concept, where the output of one command is piped or redirected to be used as input of the next command. In case of MongoDB, multiple operators are combined into a single pipeline that is responsible for processing a stream of documents. Some operators, such as $match, $limit and $skip take a document as input and output the same document in case a certain set of criteria’s is met. Other operators, such as $project and $unwind take a single document as input and reshape that document or emit multiple documents based upon a certain projection. The $group operator finally, takes multiple documents as input and groups them into a single document by aggregating the relevant values. Expressions can be used within some of these operators to calculate new values or execute string operations.
Multiple operators are combined into a single pipeline that is applied upon a list of documents. The pipeline itself is executed as a MongoDB Command, resulting in single MongoDB document that contains an array of all documents that came out at end of the pipeline. The next paragraph details the refactoring of the molecular similarities algorithm as a pipeline of operators. Make sure to (re)read the previous two articles to fully grasp the implementation logic.
2. Molecular Similarity Pipeline
When applying a pipeline upon a certain collection, all documents contained within this collection are given as input to the first operator. It’s considered best practice to filter this list as quickly as possible to limit the number of total documents that are passed through the pipeline. In our case, this means filtering out all document that will never be able to satisfy the target Tanimoto coefficient. Hence, as a first step, we match all documents for which the fingerprint count is within a certain threshold. If we target a Tanimoto coefficient of 0.8 with a target compound containing 40 unique fingerprints, the $match operator look as follows:
Only compounds that have a fingerprint count between 32 and 50 will be streamed to the next pipeline operator. To perform this filtering, the $match operator is able to use the index that we have defined for the fingerprint_count property. For computing the Tanimoto coefficient, we need to calculate the number of shared fingerprints between a certain input compound and the compound we are targeting. In order to be able to work at the fingerprint level, we use the $unwind operator. $unwind peels off the elements of an array one by one, returning a stream of documents where the specified array is replaced by one of its elements. In our case, we apply the $unwind upon the fingerprints property. Hence, each compound document will result in n compound documents, where n is the number of unique fingerprints contained within the compound.
In order to calculate the number of shared fingerprints, we will start off by filtering out all documents which do not have a fingerprint that is in the list of fingerprints of the target compound. For doing so, we again apply the $match operator, this time filtering on the fingerprints property, where only documents that contain a fingerprint that is in the list of target fingerprints are maintained.
As we only match fingerprints that are in the list of target fingerprints, the output can be used to count the total number of shared fingerprints. For this, we apply the $group operator on the compound_cid, though which we create a new type of document, containing the number of matching fingerprints (by summating the number of occurrences), the total number of fingerprints of the input compound and the smiles representation.
We now have all parameters in place to calculate the Tanimoto coefficient. For this we will use the $project operator which, next to copying the compound id and smiles property, also adds a new, computed property named tanimoto.
As we are only interested in compounds that have a target Tanimoto coefficient of 0.8, we apply an additional $match operator to filter out all the ones that do not reach this coefficient.
The full pipeline command can be found below.
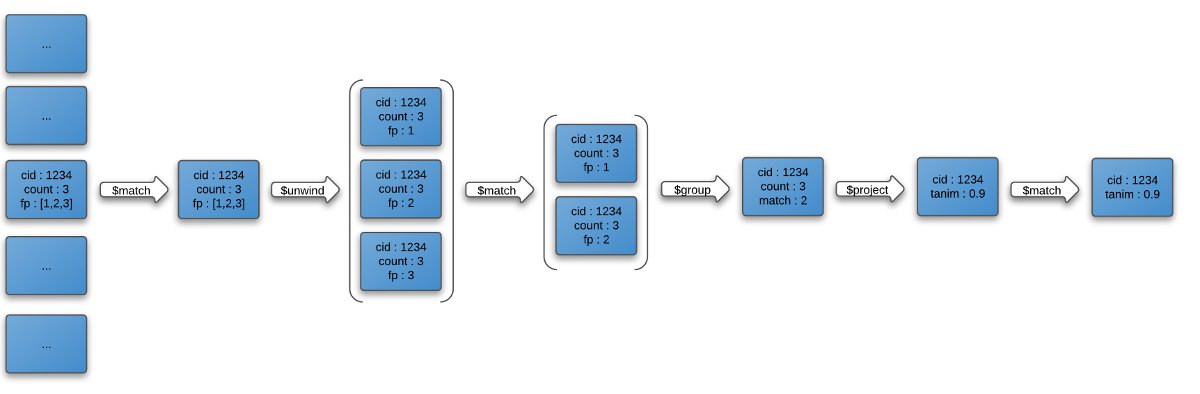
The output of this pipeline contains a list of compounds which have a Tanimoto of 0.8 or higher with respect to a particular target compound. A visual representation of this pipeline can be found below:
3. Conclusion
The new MongoDB Aggregation Framework provides a set of easy-to-use operators that allow users to express map-reduce type of algorithms in a more concise fashion. The pipeline concept beneath it offers an intuitive way of processing data. It is no surprise that this pipeline paradigm is adopted by various NoSQL approaches, including Tinkerpop’s Gremlin Framework and Neo4J’s Cypher implementation.
Performance wise, the pipeline solution is a major improvement upon the map-reduce implementation. The employed operators are natively supported by the MongoDB platform, which results in a huge performance improvement with respect to interpreted Javascript. As the Aggregation Framework is also able to work in a sharded environment, it easily beats the performance of my initial implementation, especially when the number of input compounds is high and the target Tanimoto coefficient is low. Great work from the MongoDB team!